View on GitHub
Open this notebook in GitHub to run it yourself
Prerequisites
The model uses Classiq libraries in addition to basic Python tools:Defining the Optimization Problem
Following the paper [2], create a Python Pyomo model to describe an optimization problem where a paramatrized cost expression characterizes the geometrical configuration of a protein (amino acid sequence). Without going into too many details, the paper places the protein on the grid of a tetrahedral lattice. Each amino acid can be located in any vertex of the lattice, and an index is set to each amino acid. Since each vertex has four neighbors, after locating an index in space, the next location is set by pointing in one of the four directions indicated by an integer . To ascribe a direction, assign each amino acid two qubits, thus mapping each index to a direction:- - prevents two consecutive directions to fold back.
- For the interaction, define extra qubits, which determine if the interaction is “turned on”. If so, they add a negative (beneficial) energy term , which indicates the interaction between amino acids. is a nearest neighbor (NN) interaction term, relevant for distance 1 only; therefore, add a penalty for other distances.
Creating the Protein Sequence
Define the amino acid sequence as a string that is sent to thefolding_hamiltonian() function to create an optimization model for the sequence:
Solving with the Classiq platform
We go through the steps of solving the problem with the Classiq platform, using QAOA algorithm [4]. The solution is based on defining a pyomo model for the optimization problem we would like to solve.Setting Up the Classiq Problem Instance
In order to solve the Pyomo model defined above, we use theCombinatorialProblem quantum object.
Under the hood it tranlastes the Pyomo model to a quantum model of the QAOA algorithm, with a cost function translated from the Pyomo model. We can choose the number of layers for the QAOA ansatz using the argument num_layers.
Synthesizing the QAOA Circuit and Solving the Problem
We can now synthesize and view the QAOA circuit (ansatz) used to solve the optimization problem:Output:
Output:
optimize method of the CombinatorialProblem object.
For the classical optimization part of the QAOA algorithm we define the maximum number of classical iterations (maxiter) and the -parameter (quantile) for running CVaR-QAOA, an improved variation of the QAOA algorithm [3]:
Output:
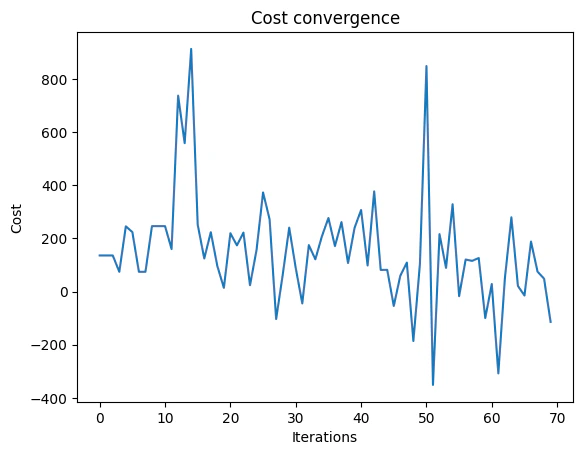
Presenting the Quantum Results
In order to get samples with the optimized parameters, we call thesample method.
Since this is a quantum solution with probablistic results, there is a defined probability for each result (shown in a histogram), where the solution is chosen as the most probable one.
Translate the solution in terms of qubits, to the location in space of the amino acids, thereby creating a sketch of the protein folding for the sequence:
| solution | probability | cost | |
|---|---|---|---|
| 14 | {‘f’: [0, 0, 1, 0, 0, 1], ‘interaction’: [1]} | 0.019531 | -2600 |
| 13 | {‘f’: [1, 1, 1, 0, 0, 1], ‘interaction’: [1]} | 0.021484 | -2600 |
| 0 | {‘f’: [1, 0, 1, 0, 0, 1], ‘interaction’: [1]} | 0.055664 | -900 |
| 114 | {‘f’: [0, 1, 1, 0, 0, 0], ‘interaction’: [1]} | 0.000488 | -900 |
| 64 | {‘f’: [0, 1, 1, 0, 0, 1], ‘interaction’: [1]} | 0.004883 | -900 |
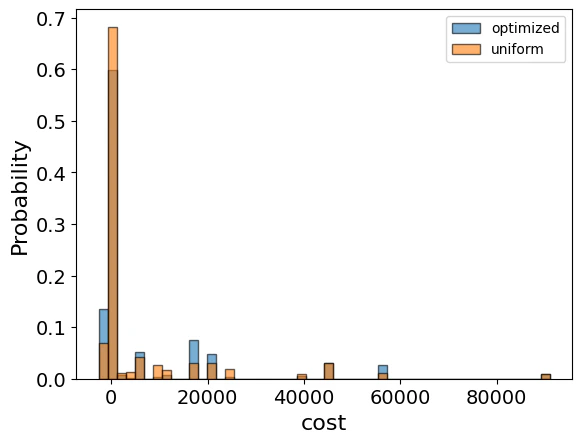
Output:
Comparing to Classical Results
Solve the optimization model for the defined amino sequence by classical optimization, and present the results, thereby comparing to the QAOA performance. A mismatch of the classical and quantum solution indicates a need to tune the QAOA parameters:Output:

